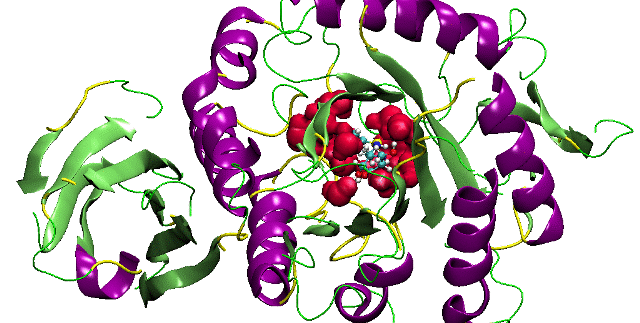
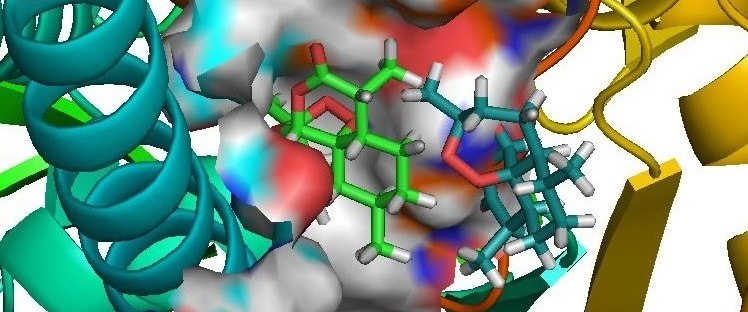
Origami of Protein Folding
Protein Folding is the ability of protein molecules to fold into their highly structured functional states defined by their amino acid sequence. The strings of amino acids that emerge from the protein synthesizing machinery, bend, loop, twist, coil and collapse on itself to produce the finished design as enzymes and other life-sustaining cellular components. The spontaneous self-assembly of protein molecules with huge numbers of degrees of freedom into a unique three dimensional structure that carries out a biological function is perhaps the simplest case of biological self-organization and one of the most remarkable achievements in biology.
Why fold proteins?
The growing genome knowledge and the pressing necessity for the discovery of new drug targets for life-threatening diseases bring the protein folding problem to the centre stage of computational molecular biology. A predictive model for protein structure remains one of the science’s holiest grails promising incredible benefits through out the medical sciences. A 3-D structure could yield invaluable insights into the function of unknown proteins, new drugable targets. Improvements in structure prediction will lead to improvement in protein design, which in turn can be directly be translated into making new enzymes, vaccines etc.
Ab initio Approach for Protein Folding
Ab initio structure prediction attempts to model proteins by starting from an extended chain and folding up the sequence on the computer. This method has an advantage that it does not depend on the existence of a previous determined structure to serve as a template. It is generally assumed that a protein sequence folds to a native confirmation or ensemble of conformations that is at or near the global free-energy minimum. Thus, the problem of finding native like conformations for a given sequence can be decomposed into two sub problems
(a) developing an efficient energy function and
(b) developing an accurate algorithm for searching the resultant energy landscape.
Many of the methods today that predict protein structure use information from the protein data bank (PDB). This information can be found in the parameters of knowledge based scoring functions, the training sets of machine learning approaches, and the coordinate libraries of methods that use fragments or templates from the PDB. In order to evaluate the accuracy of the prediction methods, Critical Assessment of Structure Prediction (CASP), a biannual, community-wide blind test of prediction methods has been conceived and implemented. In these experiments, researchers are given the amino acid sequence of target proteins and then asked to develop three-dimensional models of the final folded versions. Their predictions are compared with the actual protein structures, which has been solved experimentally by x-ray crystallography or NMR spectroscopy, but not yet published. CASP experiments have been among the most important influences in advancing this field and have provided an invaluable boost to the field. The latest of the series, CASP7 was held in November, 2006, California.
Bhageerath: Protein Tertiary Structure Prediction Server
Considered to be the holy grail of
modern biology, a solution to the protein folding problem has enormous
potentially beneficial impact on society. Prediction of three dimensional
structures of drug targets, design of biocatalysts and nanobiomachines are a few
of the multitude of foreseeable applications. At IIT Delhi, we have been
developing a computational protocol for modeling and predicting protein
structures at the atomic level. The software is named Bhageerath after the great Indian king who managed to accomplish the impossible task of getting the Ganges from heaven to earth.