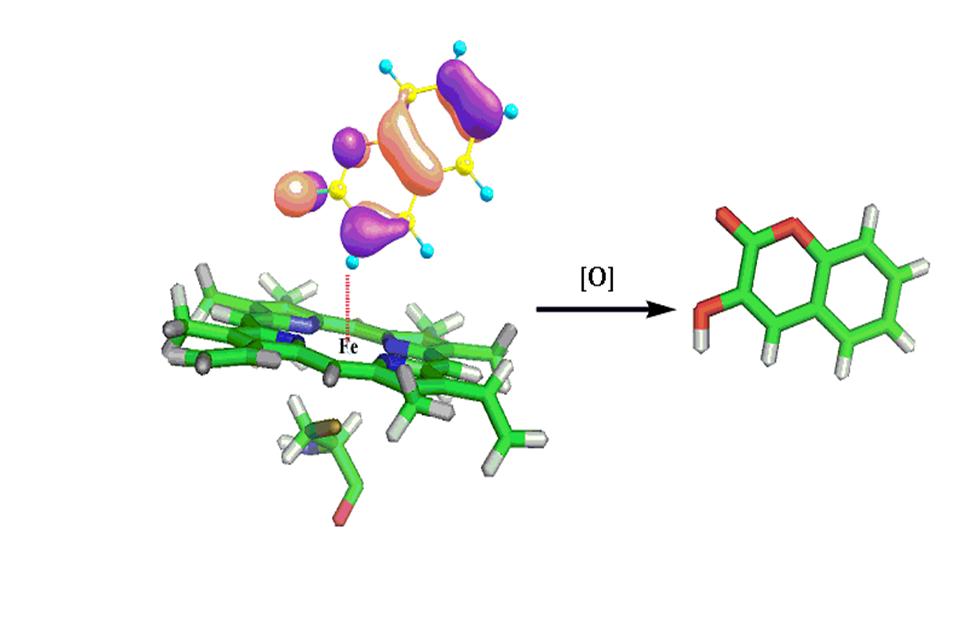
 Metabolism studies are an essential integral part of ADMET profiling of drug candidates to evaluate their safety and efficacy. Absorption and distribution can be predicated by Lipinski's rule of five. Till date there are very few in silico tools available for metabolism and toxicity prediction. Understanding metabolism in terms of metabolism sites and products (metabolites) is a central aspect in today’s drug discovery process. Knowledge of sites of metabolism (SOM) of a molecule and its biotransformation products can help not only in optimizing the lead molecule with favorable metabolic profile but also in reducing toxicity and enhancing bioavailability and bioactivity. Most of the drugs undergo metabolic transformations in the human liver which is the key source of a variety of metabolizing enzymes. Cytochrome P450s (CYPs), a superfamily of heme containing enzymes, are the major enzymes involved in drug metabolism. Cytochrome P-450 (CYP) metabolizes a wide variety of lipophilic endogenous (steroids, fatty acids) and exogenous (drugs & environmental contaminants) compounds. Binding modes of ligands with CYP decide the metabolic products. Predicting correct binding modes for a ligand with respect to any enzyme is a necessary step in structure based drug-design.
Present methodology are based on a combination of docking followed by molecular orbital (MO) calculations and knowledge based methods to predict the potential metabolic sites of a molecule and its metabolic products. This approach overcomes the limitations in finding different metabolic products of a single molecule by the same or different CYPs.
Metabolism studies are an essential integral part of ADMET profiling of drug candidates to evaluate their safety and efficacy. Absorption and distribution can be predicated by Lipinski's rule of five. Till date there are very few in silico tools available for metabolism and toxicity prediction. Understanding metabolism in terms of metabolism sites and products (metabolites) is a central aspect in today’s drug discovery process. Knowledge of sites of metabolism (SOM) of a molecule and its biotransformation products can help not only in optimizing the lead molecule with favorable metabolic profile but also in reducing toxicity and enhancing bioavailability and bioactivity. Most of the drugs undergo metabolic transformations in the human liver which is the key source of a variety of metabolizing enzymes. Cytochrome P450s (CYPs), a superfamily of heme containing enzymes, are the major enzymes involved in drug metabolism. Cytochrome P-450 (CYP) metabolizes a wide variety of lipophilic endogenous (steroids, fatty acids) and exogenous (drugs & environmental contaminants) compounds. Binding modes of ligands with CYP decide the metabolic products. Predicting correct binding modes for a ligand with respect to any enzyme is a necessary step in structure based drug-design.
Present methodology are based on a combination of docking followed by molecular orbital (MO) calculations and knowledge based methods to predict the potential metabolic sites of a molecule and its metabolic products. This approach overcomes the limitations in finding different metabolic products of a single molecule by the same or different CYPs.
 Metabolism studies are an essential integral part of ADMET profiling of drug candidates to evaluate their safety and efficacy. Absorption and distribution can be predicated by Lipinski's rule of five. Till date there are very few in silico tools available for metabolism and toxicity prediction. Understanding metabolism in terms of metabolism sites and products (metabolites) is a central aspect in today’s drug discovery process. Knowledge of sites of metabolism (SOM) of a molecule and its biotransformation products can help not only in optimizing the lead molecule with favorable metabolic profile but also in reducing toxicity and enhancing bioavailability and bioactivity. Most of the drugs undergo metabolic transformations in the human liver which is the key source of a variety of metabolizing enzymes. Cytochrome P450s (CYPs), a superfamily of heme containing enzymes, are the major enzymes involved in drug metabolism. Cytochrome P-450 (CYP) metabolizes a wide variety of lipophilic endogenous (steroids, fatty acids) and exogenous (drugs & environmental contaminants) compounds. Binding modes of ligands with CYP decide the metabolic products. Predicting correct binding modes for a ligand with respect to any enzyme is a necessary step in structure based drug-design.
Present methodology are based on a combination of docking followed by molecular orbital (MO) calculations and knowledge based methods to predict the potential metabolic sites of a molecule and its metabolic products. This approach overcomes the limitations in finding different metabolic products of a single molecule by the same or different CYPs.
Metabolism studies are an essential integral part of ADMET profiling of drug candidates to evaluate their safety and efficacy. Absorption and distribution can be predicated by Lipinski's rule of five. Till date there are very few in silico tools available for metabolism and toxicity prediction. Understanding metabolism in terms of metabolism sites and products (metabolites) is a central aspect in today’s drug discovery process. Knowledge of sites of metabolism (SOM) of a molecule and its biotransformation products can help not only in optimizing the lead molecule with favorable metabolic profile but also in reducing toxicity and enhancing bioavailability and bioactivity. Most of the drugs undergo metabolic transformations in the human liver which is the key source of a variety of metabolizing enzymes. Cytochrome P450s (CYPs), a superfamily of heme containing enzymes, are the major enzymes involved in drug metabolism. Cytochrome P-450 (CYP) metabolizes a wide variety of lipophilic endogenous (steroids, fatty acids) and exogenous (drugs & environmental contaminants) compounds. Binding modes of ligands with CYP decide the metabolic products. Predicting correct binding modes for a ligand with respect to any enzyme is a necessary step in structure based drug-design.
Present methodology are based on a combination of docking followed by molecular orbital (MO) calculations and knowledge based methods to predict the potential metabolic sites of a molecule and its metabolic products. This approach overcomes the limitations in finding different metabolic products of a single molecule by the same or different CYPs.