 |
|
|
|
|
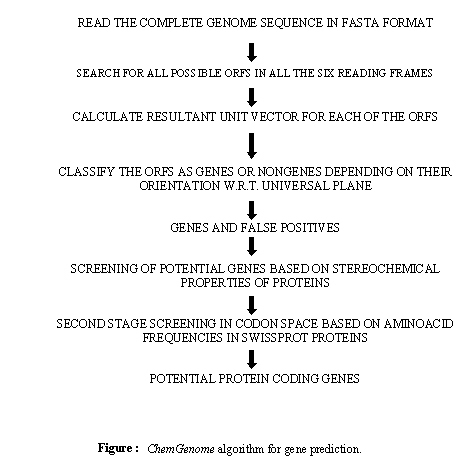
ChemGenome is an ab initio model for gene
evaluation and gene prediction in prokaryotic genomes based on
physico-chemical characteristics of codons calculated from
molecular dynamics (MD) simulations. It uses physico-chemical
properties to construct a 3D vector to answer the fundamental
question, 'What is a gene?'.
The protocol for gene
prediction has been divided into three modules that form an
automated pipeline. The methodology combines filters based on
stereochemical properties of protein sequences and frequencies
of occurrences of codons via their corresponding amino acids
from 175000 Swissprot proteins to reduce over
prediction.
The methodology has been validated on 372
prokaryotic genomes with sensitivity, specificity and
correlation coefficients averaged over 356208 genes and an
equal number of frame-shifted genes (non-genes) as 97.5%,
97.20% & 94.25% respectively.
The physico-chemical
model for gene evaluation and gene prediction for prokaryotic
genomes have been web enabled as
ChemGenome1.1 (
http://www.scfbio-iitd.res.in/chemgenome/index.jsp)
ChemGenome2.0
(http://www.scfbio-iitd.res.in/chemgenome2)
 |
| References |
[1]
Progenie "Decoding the Design Principles of Amino Acids and the Chemical Logic of Protein Sequences", Jayaram, B. Available from Nature Precedings. http://hdl.handle.net/10101/npre.2008.2135.1 2008 Read Paper
|
[2]"Prokaryotic Gene Finding based on Physicochemical Characteristics of Codons Calculated from Molecular Dynamics Simulations", Singhal P, Jayaram B, Dixit S B and Beveridge D L, Biophys. J. ,2008, 94(11), 4173-4183. [ Read Paper ]
|
[3] "A Physico-Chemical
model for analyzing DNA sequences", Dutta S, Singhal P,
Agrawal P, Tomer R, Kritee, Khurana E and Jayaram B,
J.Chem. Inf. Mod., 2006, 46(1), 78-85.[ ABSTRACT ].
|
[4] "Beyond the Wobble :
The rule of conjugates", Jayaram B, Journal of Mol.
Evol., 1997,45,704-705.
|
| |
|
In case of any Suggestions/Exceptions, Please contact
us at scfbio@scfbio-iitd.res.in | |