RM2TS: From Ramchandran Maps to
Tertiary Structures of Proteins.
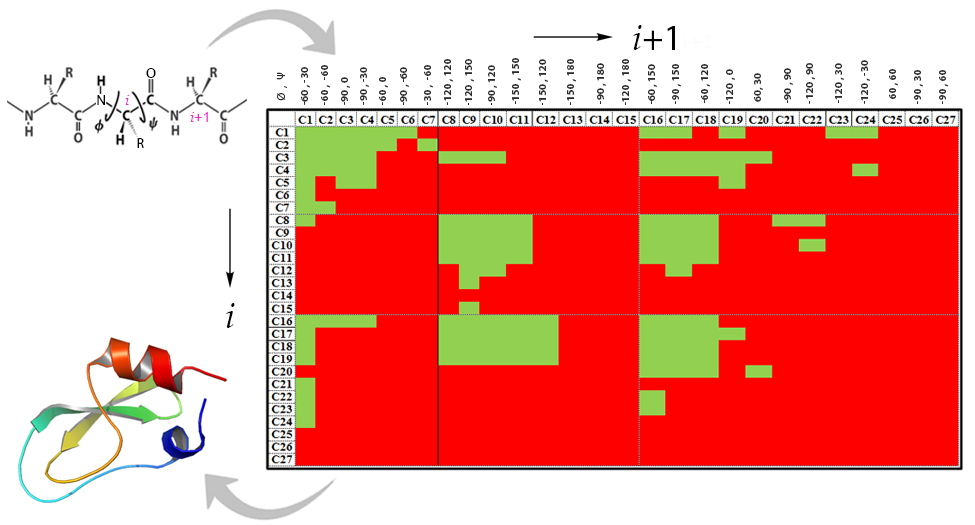
The missing link between Ramachandran map and tertiary structure

The missing link between Ramachandran map and tertiary structure
This methodology works for input sequences less than 100 amino acid residues.Please retrieve your files as the data will be deleted after 10 hours. RM2TS+ (Beta version) for bigger proteins has been developed now and is available to the users at http://www.scfbio-iitd.res.in/rm2ts+
Secondary structure prediction takes 8-10 minutes. Kindly bear with us.
Click on the classes to modify.