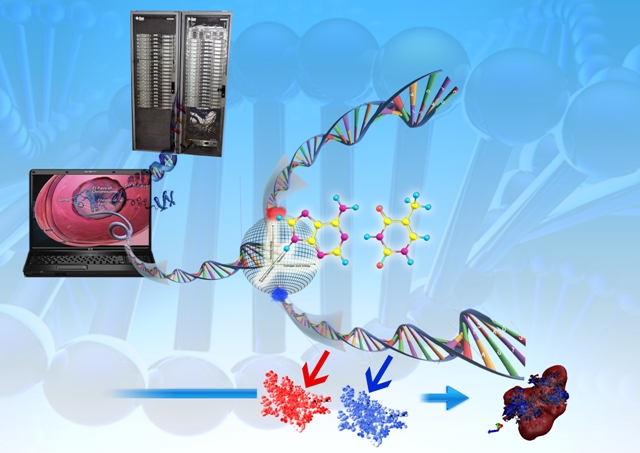
Inventus: A comprehensive drug discovery package launched by Novo Informatics Pvt Ltd (SCFBio spinoff).
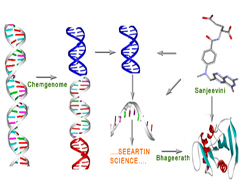
"Genome to Drug" (Dhanvantari) envisages delivering a drug molecule to society from genomic / proteomic information.
It consists of mainly three stages:
(1) interpreting the language of genomic DNA and identifying a druggable protein coding gene (Chemgenome) for a disease / disorder,
(2)
determining the three dimensional structure of the protein target (
Bhageerath) and
(3) creating a small molecule (drug) that can bind with
high affinity and specificity to the Protein/DNA target but with least toxicity to
humans (Sanjeevini).
Read more at (SCFBio Presentation 2022 Version).
To develop novel scientific methods and highly efficient algorithms, combining principles of Chemistry and Biology with Information Technology for Genome analysis, Protein structure prediction and target directed Lead molecule design pursuing the dream of SCFBio.[Reference]
Also the facility is committed towards creating a pool of Bioinformatics Professionals in the country through it's specialized training Programmes.[Reference]
The facility is providing free access of its Bioinformatics and Computational Biology tools to the global user community and free Supercomputing time to Students & Reseachers with-in the country.
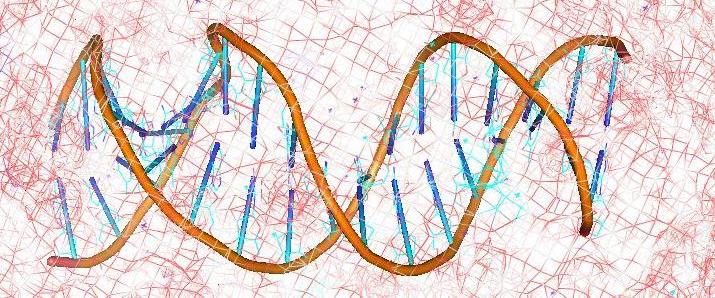
ChemGenome:Chemgenome identifies protein coding / regulatory regions
from genomic information based on the physico-chemical principles
behind genome organization.
Genome Analysis Software Suite
http://pubs.acs.org/doi/abs/10.1021/ci050119x
DOI:10.1529/biophysj.107.116392
doi:10.1093/nar/gkw1236
https://doi.org/10.1093/nar/gkab098
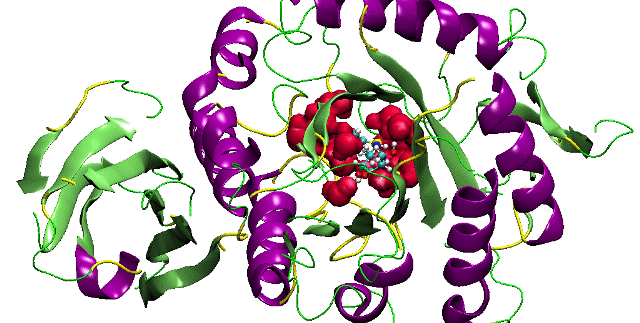
Bhageerath H:Bhageerath-H has currently
reached 60% accuracy in predicting 3D structures of soluble proteins to
within 5Ang from the native in the absence of experimental structures.
A Homology ab-intio Hybrid Web server
for Protein Tertiary Structure Prediction
10.1039/B502226F
https://doi.org/10.1093/nar/gkl7897
doi:10.1186/1471-2105-15-S16-S7
DOI: 10.1021/acs.biochem.7b01073
doi:10.1093/database/bay021
RM2TS:RM2TS is "It is a newly developed ab initio methodology which predicts the tertiary structure of a small protein ( < 100 amino acid residues) based on input amino acid sequence using higher order Ramachandran Maps.The main USP of the methodology being the short time it takes for predicting the structure ( no mail id is required to be input by the user as the protocol takes 5-10 minutes for outputting the results on screen). Even in such small time, the accuracy is not compromised on with current % around 75% for small proteins with predicted secondary structure.
From Ramchandran Maps to Tertiary Structures of Proteins.
DOI: 10.1021/acs.jpcb.5b02999
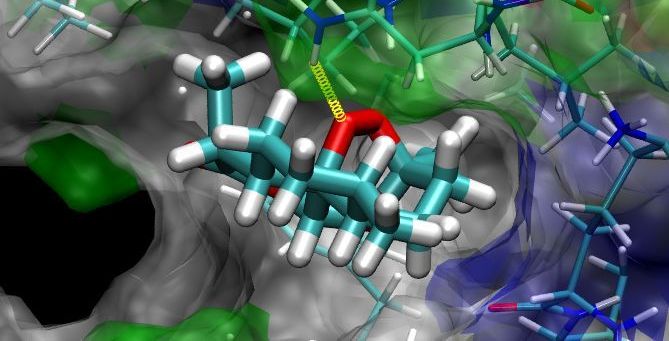
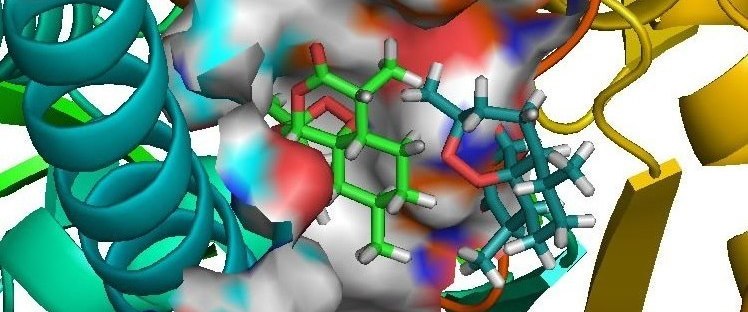
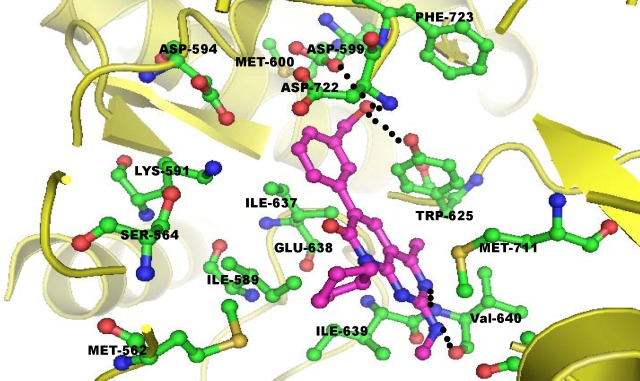
Sanjeevini:Sanjeevini helps to create lead compounds (drugs) targeted to
Protein/DNA. Sanjeevini has been
able to generate a few sub-micromolar compounds against malaria & breast
cancer.These are under further development
In-Silico Drug Design Software
http://www.niscair.res.in/sciencecommunication/research-journals/rejour/ijca/ijca2k6/ijca_jan06.asp#p21
http://www.biomedcentral.com/1471-2105/13/S17/S7
doi:10.1038/nature16963
DOI:10.1111/febs.14707
https://doi.org/10.1007/978-981-15-8936-2_1

Ruchika Bhat, Rahul Kaushik, Ankita Singh, Debarati DasGupta, Abhilash Jayaraj, Anjali Soni, Ashutosh Shandilya, Vandana Shekhar, Shashank Shekhar, B. Jayaram, " A comprehensive automated computer-aided discovery pipeline from genomes to hit molecules" Chemical Engineering Science, 2020, 222 (115711).
https://doi.org/10.1016/j.ces.2020.115711.